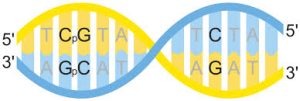
CpG characteristics
The CpG sites or CG sites are regions of DNA where a cytosine nucleotide is followed by a guanine nucleotide in the linear sequence of bases along its 5′ → 3′ direction.
CpG refers to a cytosine (C) followed by a guanine (G) in the 5′ to 3′ direction along the DNA strand, with the “p” indicating the phosphate bond between them.
These regions are called “islands” because they stand out from the rest of the genome due to their distinct properties.
CpG sites occur with high frequency in genomic regions called CpG islands (or CG islands).
CpG islands are regions of DNA that contain a high frequency of CpG, cytosine followed by guanine, dinucleotides.
These regions are typically found in the promoter regions of genes and are often associated with gene regulation.
CpG islands are usually unmethylated in normal cells, which allows for the binding of transcription factors and the initiation of gene expression.
Methylation of CpG islands can lead to gene silencing or reduced gene expression.
Abnormal DNA methylation patterns of CpG islands have been implicated in various diseases, including cancer and developmental disorders.
CpG islands are regions in DNA sequences that are characterized by a high frequency of CpG dinucleotides.
CpG islands have a higher-than-expected frequency of CG dinucleotides compared to the rest of the genome.
CpG islands are often found near the transcription start sites or promoters of genes, especially housekeeping genes and many tissue-specific genes.
In contrast to most CpG sites in the genome, which are typically methylated, meaning a methyl group is added to the cytosine, CpG islands are usually unmethylated under normal conditions.
The methylation status of CpG islands plays a crucial role in gene regulation.
When a CpG island becomes methylated, it often leads to the silencing of the associated gene.
CpG islands are important epigenetic markers, as changes in their methylation patterns can occur during development, in response to environmental factors, or in diseases like cancer.
Some CpG islands are involved in genomic imprinting, where genes are expressed differently depending on whether they are inherited from the mother or father.
In females, CpG island methylation is involved in the inactivation of one X chromosome.
A region is typically considered a CpG island if it is at least 200 base pairs long, has a G+C content greater than 50%, and an observed/expected CpG ratio greater than 0.6.
In cancers, there is a global hypomethylation of the genome, but localized hypermethylation of CpG islands near tumor suppressor genes, which can lead to their silencing and contribute to tumorigenesis.
Understanding CpG islands is crucial for comprehending gene regulation, epigenetics, developmental biology, and the molecular basis of diseases like cancer.
Hypermethylation of CpG islands in promoter regions can result in the inactivation of tumor suppressor genes or other genes involved in regulating cell growth, leading to uncontrolled cell proliferation and the development of cancer.
70% to 80% of CpG cytosines are methylated.
Methylating the cytosine within a gene can change its expression, a mechanism that is part of a larger field of science studying gene regulation that is called epigenetics.
Methylated cytosines often mutate to thymines.
About 70% of promoters located near the transcription start of the site of a gene, proximal promoters, and contain a CpG island.
CpG is shorthand for 5’—C—phosphate—G—3′ , that is, cytosine and guanine separated by only one phosphate group; phosphate links any two nucleosides together in DNA.
The frequency of CpG dinucleotides in human genomes is less than one-fifth of the expected frequency.
CpG islands are typically 300–3,000 base pairs in length, and have been found in or near approximately 40% of promoters of mammalian genes.
Over 60% of human genes and almost all house-keeping genes have their promoters embedded in CpG islands.
CpG islands typically occur at or near the transcription start site of genes, particularly housekeeping genes.
About 70% of promoters located near the transcription start site of a gene (proximal promoters) contain a CpG island.
Distal promoter elements also frequently contain CpG islands.
CpG islands also occur frequently in promoters for functional noncoding RNAs such as microRNAs.
Methylation of CpG islands silences genes.
In cancers, loss of expression of genes occurs about 10 times more frequently by hypermethylation of promoter CpG islands than by mutations.
Hypomethylation of CpG islands in promoters results in overexpression of the genes or gene sets affected.
Specific genes with colon cancer are associated with hypermethylated promoters.
MicroRNAs whose promoters are hypermethylated in colon cancers are at frequencies between 50% and 100% of cancers.
MicroRNAs (miRNAs) are small endogenous RNAs that pair with sequences in messenger RNAs to direct post-transcriptional repression.
Each microRNA represses several hundred target genes.
Thus microRNAs with hypermethylated promoters may be allowing over-expression of hundreds to thousands of genes in a cancer.
In cancers, promoter CpG hyper/hypo-methylation of genes and of microRNAs causes loss of expression or sometimes increased expression of far more genes than does mutation.
DNA repair genes with hyper/hypo-methylated promoters in cancers
DNA repair genes are frequently repressed in cancers due to hypermethylation of CpG islands within their promoters.
Many types of cancer are deficient in one or more DNA repair genes due to hypermethylation of their promoters.
Promoter hypermethylation of the DNA repair gene MGMT occurs in 93% of bladder cancers, 88% of stomach cancers, 74% of thyroid cancers, 40%-90% of colorectal cancers and 50% of brain cancers,
Promoters of genes, PARP1 and FEN1, are hypomethylated and these genes were over-expressed in numerous cancers.
PARP1 and FEN1 are essential genes in the error-prone and mutagenic DNA repair pathway.
PARP1 is over-expressed in tyrosine kinase-activated leukemias, in neuroblastoma, in testicular and other germ cell tumors, and in Ewing’s sarcoma.
FEN1 is over-expressed in the majority of cancers of the breast,prostate, stomach, neuroblastomas, pancreatic,and lung.
DNA damage appears to be the primary underlying cause of cancer.
If DNA repair is deficient, DNA damages tend to accumulate.
Sudamage can increase mutational errors during DNA replication due to error-prone translesion synthesis.
Excess DNA damage can also increase epigenetic alterations due to errors during DNA repair.
CpG island hyper/hypo-methylation in the promoters of DNA repair genes are likely central to progression to cancer.
Age has a strong effect on DNA methylation levels on tens of thousands of CpG sites.
There is an accurate biological or epigenetic clock or DNA methylation age in humans.
DNA demethylation of CpG sites during memory formation depends on initiation by ROS.
ROS-dependent demethylation of CpG sites in gene promoters within neuron DNA is central to memory formation.
